A crew of researchers from Google Analysis and the College of California, Santa Cruz has launched DeepSomatic, an AI mannequin that identifies genetic mutations in most cancers cells. The research with Kids’s Mercy discovered 10 mutations in childhood leukemia cells that have been missed by different instruments. DeepSomatic has a most cancers genome somatic small variant caller that works throughout Illumina quick reads, PacBio HiFi lengthy reads, and Oxford Nanopore lengthy reads. This technique extends DeepVariant to detect single nucleotide variants and small insertions and deletions in whole-genome and whole-exome information, and helps regular tumor and tumor-only workflows, together with FFPE fashions.
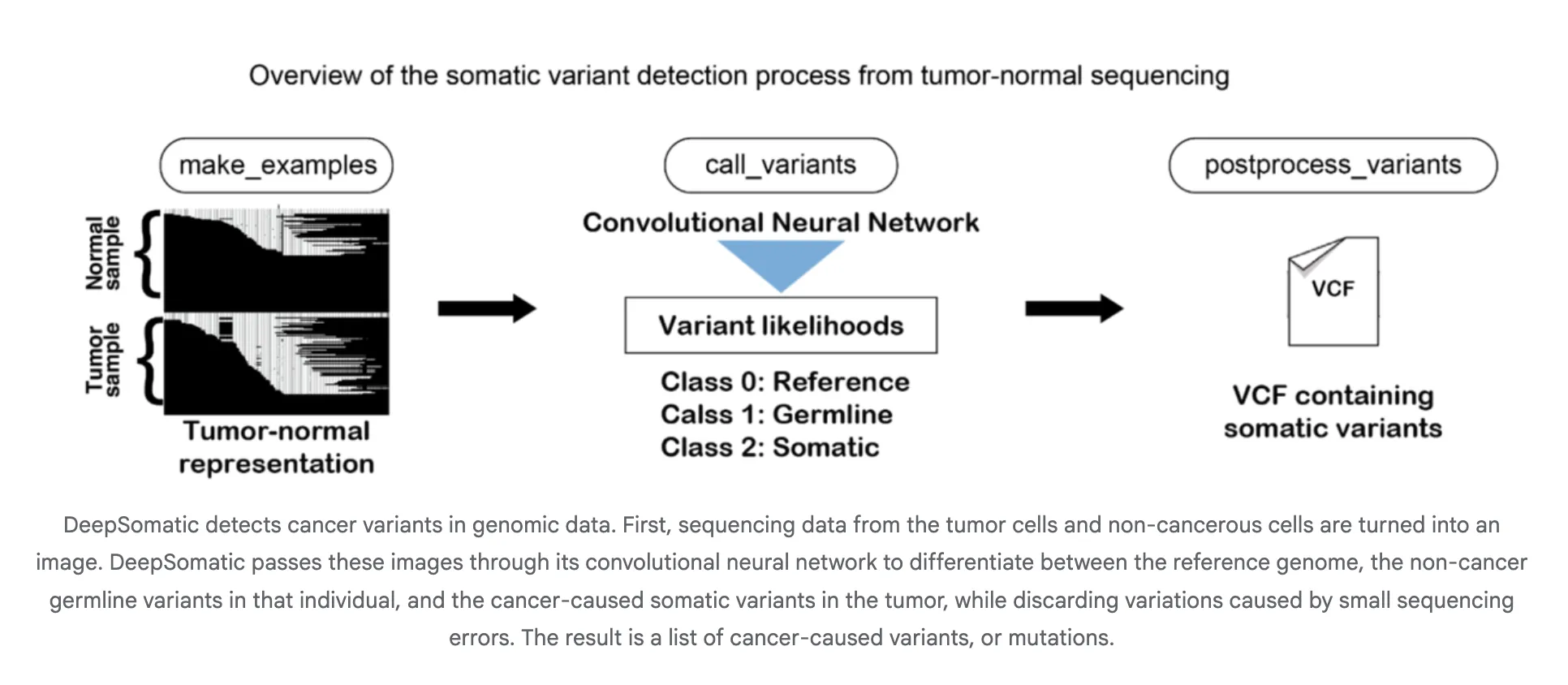
How does it work?
DeepSomatic transforms aligned reads into tensor-like photographs that encode pileup, base high quality, and alignment context. The convolutional neural community classifies the candidate web site as somatic or not, and the pipeline outputs a VCF or gVCF. This design is platform-independent, as tensors summarize native haplotypes and error patterns throughout applied sciences. Google researchers describe their method as specializing in distinguishing between inherited and purchased variants, together with troublesome samples similar to glioblastoma and childhood leukemia.
Datasets and benchmarks
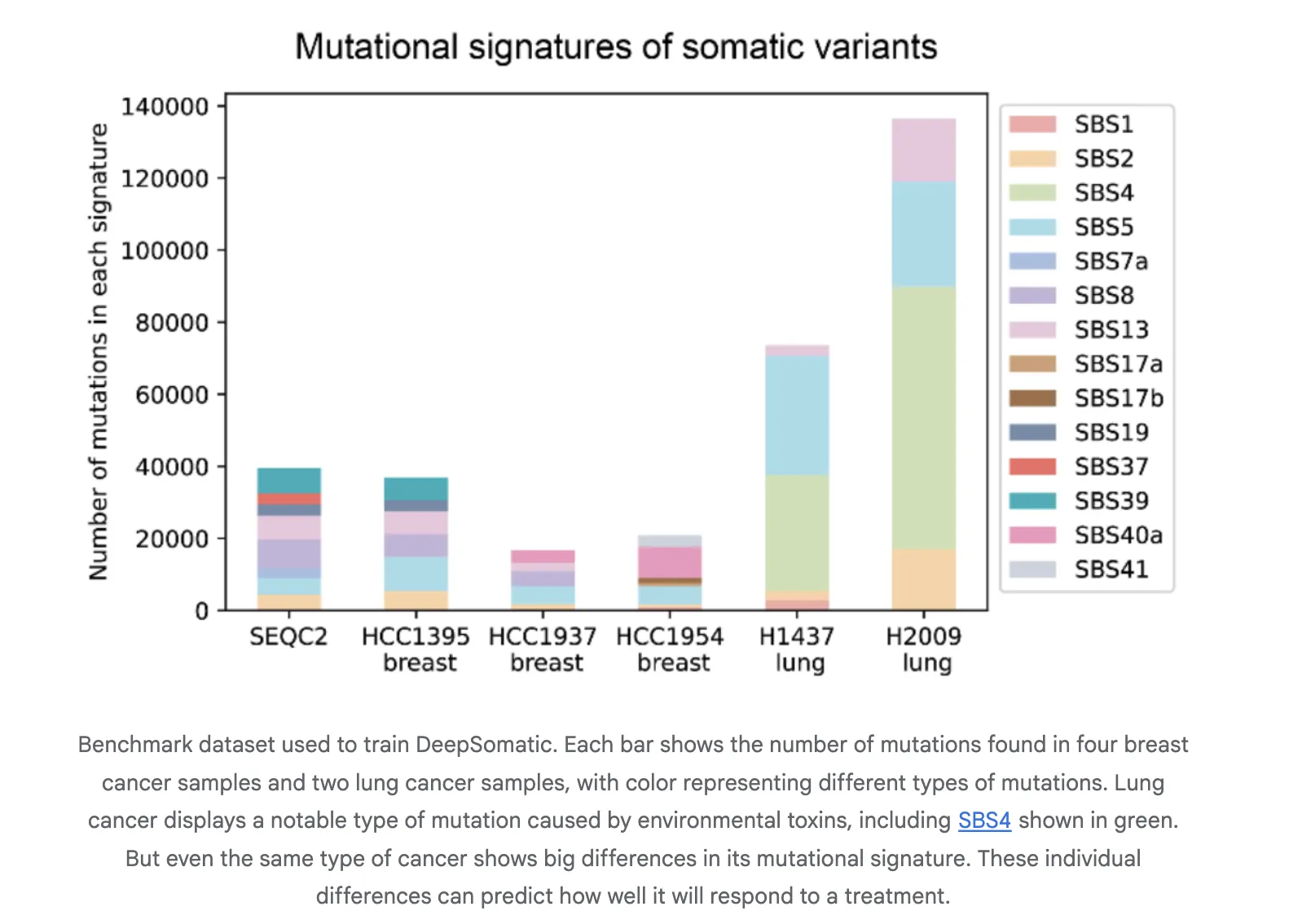
We use CASTLE, Most cancers Requirements Lengthy learn evaluation, for coaching and analysis. CASTLE comprises six matched tumor and regular cell line pairs sequenced complete genome on Illumina, PacBio HiFi, and Oxford Nanopore. The analysis crew releases benchmark units and accessions for reuse. This bridges the hole in multi-technology bodily coaching and testing sources.
Reported outcomes
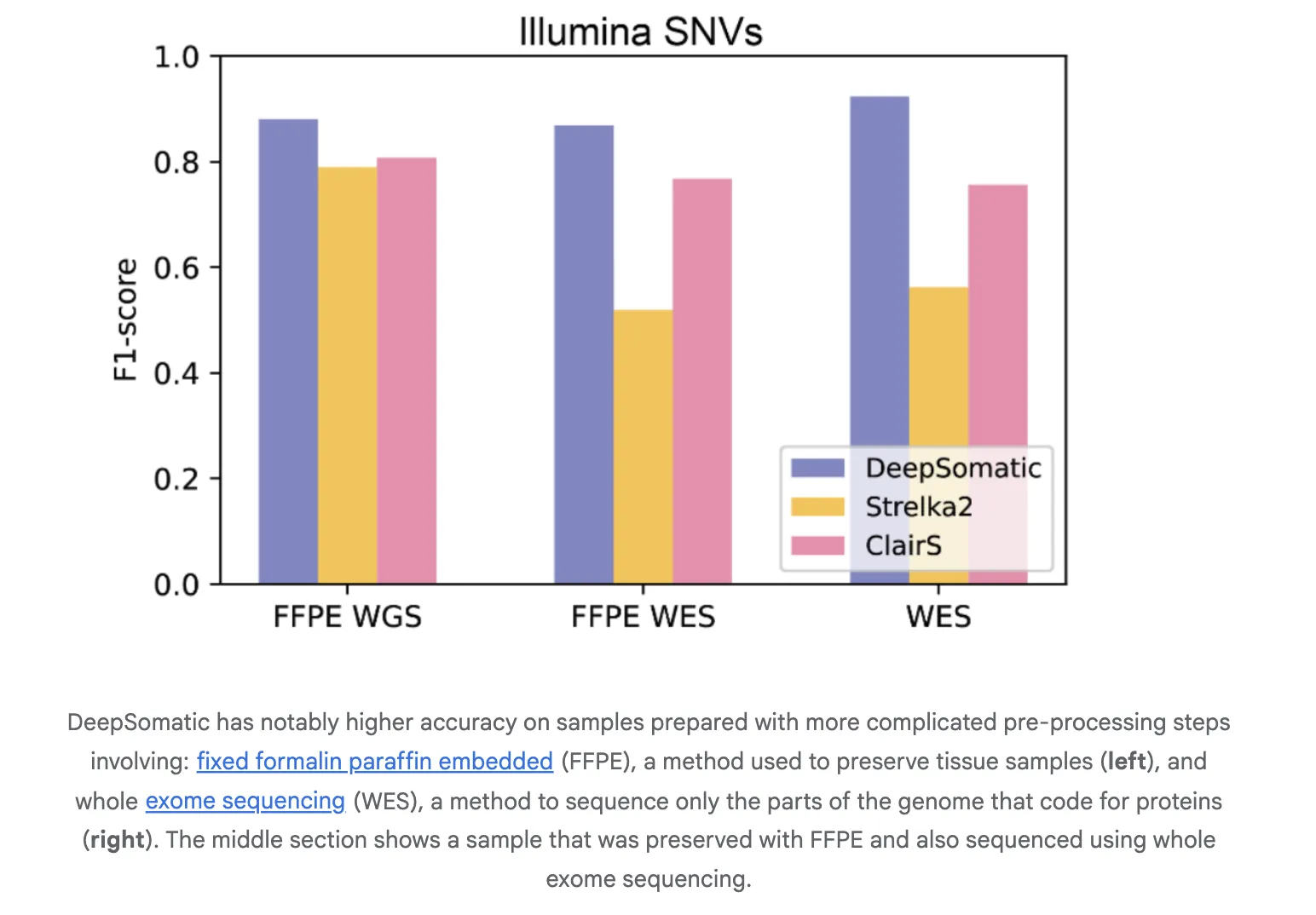
The analysis crew reported constant advantages over extensively used strategies for each single nucleotide variants and indels. For Illumina indels, the subsequent greatest strategies are F1 at about 80 % and DeepSomatic at about 90 %. For PacBio indels, the subsequent greatest technique is lower than 50 %, and DeepSomatic is greater than 80 %. The baseline contains SomaticSniper, MuTect2, and Strelka2 for brief reads and ClairS for lengthy reads. This research experiences 329,011 somatic mutations throughout the baseline and extra archival samples. The Google analysis crew experiences that DeepSomatic is especially immune to indels and performs higher than present strategies.
Generalization to actual samples
The analysis crew is evaluating most cancers metastasis past the coaching set. Glioblastoma samples present restoration of identified drivers. Childhood leukemia samples are examined in tumor-only mode the place clear regular values should not accessible. This device recovers identified calls and experiences extra variants inside that cohort. These research present that the illustration and coaching scheme generalizes to new illness conditions and settings with out matching normals.
Essential factors
DeepSomatic detects somatic SNVs (single nucleotide variants) and indels throughout Illumina, PacBio HiFi, and Oxford Nanopore and is predicated on DeepVariant methodology. This pipeline helps regular tumor and tumor-only workflows, contains FFPE WGS and WES fashions, and is launched on GitHub. Encode the learn pileup as a tensor-like picture, classify the somatic areas utilizing a convolutional neural community, and output a VCF or gVCF. The CASTLE dataset containing six matched tumor-normal cell line pairs sequenced on three platforms is used for coaching and analysis, offering benchmarks and accessions. Reported outcomes present that roughly 90% indel F1 in Illumina and greater than 80% indel F1 in PacBio exceeded the frequent baseline, with 329,011 somatic mutations recognized throughout reference samples.
DeepSomatic is a sensible step to calling somatic variants throughout sequencing platforms, and the mannequin retains DeepVariant’s picture tensor illustration and convolutional neural community, permitting the identical structure to increase from Illumina to PacBio HiFi to Oxford Nanopore with constant preprocessing and output. The CASTLE dataset is the fitting selection, offering matched tumor and regular cell traces throughout three applied sciences to reinforce coaching and benchmarking and enhance reproducibility. The reported outcomes spotlight indel detection accuracy of roughly 90% F1 for Illumina and >80% for PacBio in opposition to low baselines, which addresses a long-standing weak spot in indel detection. This pipeline helps WGS and WES, regular tumor and tumor solely, and FFPE to match real-world laboratory constraints.
Take a look at technical papers, technical particulars, datasets, and GitHub repositories. Be at liberty to go to our GitHub web page for tutorials, code, and notebooks. Additionally, be at liberty to comply with us on Twitter. Additionally, do not forget to affix the 100,000+ ML SubReddit and subscribe to our e-newsletter. cling on! Are you on telegram? Now you can additionally take part by telegram.

Michal Sutter is a knowledge science skilled with a grasp’s diploma in information science from the College of Padova. With a powerful basis in statistical evaluation, machine studying, and information engineering, Michal excels at reworking advanced datasets into actionable insights.
🙌 Observe MARKTECHPOST: Add us as your most well-liked supply on Google.






